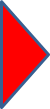

r.randomwalk is a flexible and multi-functional conceptual tool for backward- and forward-analyses of mass movement propagation. Mass points are routed from defined release raster cells of one to many mass movements through a digital elevation model until a defined break criterion is reached. Lateral spreading is ensured by a constrained random walk approach. Compared to existing tools, the major innovative features of r.randomwalk are:
- multiple break criteria can be combined to compute an impact indicator score;
- the uncertainties of break criteria can be considered by performing multiple parallel computations with randomized parameter settings, resulting in an impact indicator index in the range 0-1;
- the results can be evaluated against observations and visualized (built-in functions to do so are available);
- observed landslides can be back-analyzed to derive the density distribution of the observed angles of reach. This distribution can be employed to compute an impact probability for each raster cell.
Further, impact indicator scores and probabilities can be combined with release indicator scores or probabilities. Risk indicator scores can be computed if possible objects at risk are given.
r.randomwalk offers the possibility for exploiting the advantages of multi-core computers in order to reduce computational time and allows choosing inhowfar to use the GRASS Segment Library. It contains comprehensive functionalities for parameter optimization and model avaluation, based on reference observations, and for the visualization of the results.

Requirements
r.randomwalk 2.1 runs on LINUX operating systems. It relies on GRASS 7 or 8, Python 3, and R.
r.randomwalk 2.1 was developed and tested on Ubuntu 20.04 LTS and 24.04LTS, but is expected to work in other LINUX environments, too. It relies on GRASS GIS 7 or GRASS 8 and Python 3. Installation of r.randomwalk 2.1 requires the grass-dev package. Visualization and evaluation of the model results (flag v) makes use of the R Project for Statistical Computing. The following packages of R are required in order to fully explore the functionalities offered by r.randomwalk 2.1: sp, ROCR, raster, and terra. The code builds on Python and C. The script grass.install.sh provided with r.randomwalk 2.1 can be used to automatically install all the software needed on fresh installations of Ubuntu, and probably also on installations of other LINUX environments. However, please note that LINUX environments and the related software are dynamically evolving, so that the script might be outdated quickly, requiring manual modifications in case of failure. Manual interventions might also be necessary if older installations of some of the required software do exist on the system. The script has to be called from the terminal as superuser (requiring administrator rights): sudo sh grass.install.sh.

Installation
r.randomwalk 2.1 can be easily installed through the command g.extension.
As soon as all software requirements are fulfilled, the installation of r.randomwalk 2.1 is straightforward, using the built-in GRASS module g.extension. Assuming that the folder with the r.randomwalk source code is stored in the directory /home/user1/, installation just requires the execution of the following command:
g.extension extension=r.randomwalk url=/home/user1/r.randomwalk/

Operation
r.randomwalk 2.1 is most efficiently operated through shell scripts.
r.randomwalk 2.1 is most efficiently operated through shell scripting. Even though beginners often feel uncomfortable first in working with the terminal and with shell scripts, most of them quickly understand that this is a highly comfortable and efficient way to launch simulations and to analyze the results in a systematic way.
First, you have to launch GRASS with the desired Location and Mapset. Please consult the official GRASS GIS website to learn how to do so. You will not use the GUI of GRASS, but work in the terminal. The command line prompt should start with the GRASS version and, in brackets, the location. This means that you are really in GRASS. You can now start r.randomwalk 2.1 by typing the arguments directly into the command line:
r.randomwalk -q -v prefix=test elevation=test_elev releasefile=test_releasefile.txt models=1,1,4,-9999,-9999 mparams=1,500,500,100,20,5,2
In this case, r.randomwalk 2.1 would route one or more landslides defined in the text file test_releasefile.txt down throught the elevation map test_elev, according to the rules defined through the options model and mparams, compute an impact indicator score, and produce a set of map layouts. The results would be collected in a folder named test_results, which would be stored in the current directory. The input raster map test_elev would have to be available as a GRASS raster in the current mapset.
You can learn much more about the input of r.randomwalk in the section below. However, also with this minimum input it can be demonstrated that this way of operation is rather efficient: if you wish to run the simulation again, you can just recover the string by typing the upward key.
But we still go one step further. We can also write the command we have just typed into the terminal in an executable text file, and instead execute this file. Whereas this does not look very straightforward on a first glance, it dramatically increases our flexibility: we can now write several commands into the same script - for example, running r.randomwalk for several times with different input. We can go for a beer, to sleep, or even on holiday, and see the results when we have returned, without any intervention required in between.
It is recommended to create a folder for each project we are working on. So, let us create the folder projects/test in our home directory. Within this folder, let us create a text file with the name test.randomwalk.start.sh. The suffix sh stands for shell script, which is the most common type of executable file in LINUX environments. We now copy the text we have executed from the command line into this script, and save our changes. Then, back in the terminal, we have to change to the directory where the script is stored, and execute the script by typing sh and its name:
cd projects/test/
sh test.randomwalk.start.sh
You will find the folder with the results of the computation in the directory projects/test. Using shell scripts, you cannot only run several calculations at a time. They also help you to:
- Import tiff or ascii rasters into GRASS (command r.in.gdal)
- Create generic landcapes (command r.mapcalc)
- Preprocess raster maps (command r.mapcalc)
Examples of such start scipts can be found in the training data.

Input
r.randomwalk 2.1 is very flexible with regard to input.
A large choice of options is available for the definition of the initial conditions and the parameterization of r.randomwalk 2.1. Not all options are needed for all types of calculations - in fact, simple calculations can be performed with very little input. You can filter the set of options by clicking on one of the keywords below. Klicking on further keywords constrains the number of options. The entire set can be recovered by clicking on All.
All
Key parameters
Operational parameters
Release parameters
Multi-core processing
Back-calculation
Impact probability
Impact ind. score
Connectivity score
Risk estimation
Integrated approach
Calibration and evaluation
These options are important for setting the scene of the simulation.
These are the options beginners should start with.
These options serve for parameter calibration and for the evaluation of the model results.
This set of options can be used to define landslide release.
This set of options is relevant for the back-calculation of travel distances and angles of reach of observed landslides.
These options serve for the input for calculations of the impact indicator score.
This set of options is relevant for the computation of impact probability.
This set of options can be used to simulate geomorphological connectivity.
This set of options is relevant for multi-core processing, resulting in an impact indicator index.
These options support the simulation of the integrated landslide probability, using the output of the r.landslides.statistics model (see Mergili et al., 2019).
This set of options can be used to produce a risk indicator score, based on the overlay of the impact indicator score and a layer of potential objects at risk.

-a
Average of impact probability
This flag is only available in combination with the flag p. In contrast to the default, where the maximum out of the impact probabilities of overlapping cases is used as the overall impact probability, the average out of the impact probabilities of overlapping cases is applied.

-b
Back-calculate observations
This mode is useful to explore the angle of reach and the maximum travel distance of observed mass movements. Instead of applying the break criteria set in the option models, each random walk stops when leaving the impact area of the case (option impactarea).

-c
Compute connectivity
With this flag a connectivity score is computed, describing the degree to which a release point or a release raster cell is connected to those raster cells which are larger than 0 in the option impactobjects, which is mandatory with flag c.

-k
Keep result GRASS raster maps
Particularly with a very large number of model runs (flag m and option cores), a huge amount of GRASS raster maps is produced which are often not needed, but require a considerable amount of disk space. Without the flag k all result GRASS raster maps stored in the active mapset are deleted, with the flag they are kept.

-m
Multiple model runs with random parameters
Multiple model runs are executed with parameters varied in a random or controlled manner (option sampling) between upper and lower boundaries given by models and mparams. The results are combined to an impact indicator score map. Further, input and output parameter tables suitable as input for the sensitivity analysis and parameter optimization tool AIMEC (Fischer, 2013) are created.

-n
Multiple model runs with random release
Multiple model runs are executed with random subsets of the releasefile or the releasemap. This flag is useful in combination with the flag p in order to clearly separate between a set of cases used for model development and another set employed for evaluation. Separate cumulative density functions are derived for each subset.

-p
Probabilities
An impact probability map - and, if release probabilities map (through the option probmap or in the releasefile) are given, also a composite probability map - is computed.

-q
Scores
An impact indicator score map - and, if release indicator scores (through the option scoremap or in the releasefile) are given, also a composite indicator score map - is computed.

-s
Separate random walks for each model
The parameters Lctrl, Lseg, Rmax, fbeta and fdir (see options models and mparams) are defined separately for each model. Activating this flag increases the flexibility of the analysis, but - as the random walks have to be run separately for each model - also the computational time.

-v
Evaluation and visualization of model results
ROC Plots are generated, evaluating the impact indicator index (flag m) or impact probability maps (flag p) against the observed deposition areas (option depositmap). Map layouts of the main results are produced. In the back-calculation mode (flag b), cumulative density plots are created. A table evaluating the prediction sucess of all model runs is generated with the flag m.

-x
Start routing from raster cells
Random walks start from all raster cells where releasemap>0 or, if a release probability map (option probmap, with the flag p) or a release indicator score map (option scoremap, with the flag q) is given, from all raster cells in the area of interest. If a releasefile, but none of the options probmap or scoremap is given, the release probability (with the flag p) or release indicator score (with the flag q) is taken from the releasefile.

prefix
Prefix for output files and folders
Mandatory prefix applied to all output files and folders. Any type of string can be used, but it is recommended to choose a combination of 3-6 characters to shortly describe the simulation.

cores
Number of cores
cores=8
When performing multiple model runs (flags m or n), the number of processors to be used is a mandatory input. E.g., cores=8 induces the use of 8 processors.

cellsize
Raster cell size in metres
cellsize=[cell size of elevation raster map]
raster cell size in metres to be used for the model. If the raster cell size is not given, it is taken from the input elevation raster map (option elevation).

aoicoords
Coordinates delineating the area of interest
Series of four coordinates (N, S, E, W) delineating the bounding rectangle to applied for the analysis. If this parameter is not given, the bounding rectangle will be determined from the elevation raster map (option elevation).

aoimap
Name of raster map delineating the area of interest
Name of integer input raster map of the area of interest to be considered in the model. All raster cells with values > 0 are interpreted as part of the area of interest, all other raster cells are excluded from the computation. If no area of interest map is given, all non-no data raster cells of the elevation raster map (option elevation) are used. The bounding box defined by aoicoords applies in any case.

elevation
Name of elevation raster map
Name of input elevation raster map. The unit is metres.

releasefile
Name of text file with release information
The relative path to the text file defining the cases to be routed is an optional input. Alternatively, the random walks may start from a releasemap (flag x), a release probability map (option probmap, flags p and x) or a release score map (option scoremap, flags q and x).
Each line of the case file characterizes on specific case by ten parameters, separated by tabulators:
- ID: Case id (a positive integer number, it is not necessary to keep the ids in ascending order);
- TYPE: Case type (an integer number denoting the type of the case according to a user-defined scheme);
- M: Case magnitude or -9999 for no data;
- Qp: Case peak discharge in cubic metres per second or -9999 for no data;
- RIS: Release indicator score associated to the case or -9999 for no data;
- Pr: Release probability associated to the case or -9999 for no data;
- Xr, Yr: y and y coordinates of the release point (the reference point for all geometric break criteria);
- Xs, Ys: x and y coordinates of the start point (the point where routing starts, it may be identical to or downslope from the release point).
This file requires a header line following the above naming conventions. The following example shows the possible content of a release file for two cases where the magnitudes are expressed by release volumes and release indicator scores are given.
| ID | TYPE | V | QP | RIS | PR | XR | YR | XS | YS |
| 37 | 2 | 27350 | -9999 | 1 | -9999 | 250222 | 327860 | 250336 | 327522 |
| 44 | 1 | -9999 | 8500 | 4 | -9999 | 261570 | 338126 | 261137 | 337980 |

caserules
Assignment of models to cases
This optional parameter defines which models (see option models) are applied to which types of cases (see option releasefile). For each type of cases, the integer value denoting the type is followed by as many comma-separated integers as models are given in the option models. 1 means that the corresponding model shall be applied to the case of the corresponding type, 0 means that this model shall not be used. For two models and two cases, the entry could look as follows:
caserules=1,0,1,2,1,1
If caserules is not given, all models are applied to all cases.

releasemap
Name of raster map of release areas
Name of an optional input integer raster map defining the observed release area of each case. The raster cell values have to correspond to the case id given in the releasefile. Zero stands for no case. With the flag x, random walks are started from all raster cells of all release areas. If a release probability map (option probmap, flags p and x) or a release score map (option scoremap, flag q) is provided, the releasemap is not used for the computation, but only for visualization (flag v).

depositmap
Name of raster map of observed deposition areas
Name of an optional input integer raster map defining observed deposition area of each case. Positive raster cell values define areas with an observed deposition, raster cell values of zero define areas without observed deposition. This map is needed for evaluation and visualization only (flag v). For evaluation purposes it is recommended to set the part of the impact area (option impactmap) not defined as deposition area to no data.

impactmap
Name of raster map of observed impact areas
Name of an optional input integer raster map defining the observed impact area of each case. When used for back-calculation (flag b), the raster cell values have to correspond to the case id given in the releasefile. zero stands for no case. When used for evaluation and/or visualization only (flag v), areas with observed impact should be indicated by positive values, areas with no observed impact by zero.

probmap
Name of release probability raster map
Name of optional input raster map of the raster cell-based release probability Pr in the range 0-1, applicable with the flag p.

scoremap
Name of raster map indicating release scores
Name of optional integer input raster map of the raster cell-based release indicator score RIS in the range 0-6, applicable with the flag q.

impactobjects
Name of raster map of possible objects at risk
Optional raster map where raster cells coinciding with defined impact objects such as buildings take values larger than zero, and raster cells not coinciding with such objects take values of zero.

objectscores
Name of text file characterizing possible objects at risk
The relative path to the text file characterizing the objects denoted by impactobjects is an optional input. Each line of the file represents one object, characterized by the following parameters:
- Object id (a positive integer number, it is not necessary to keep the ids in ascending order);
- Object type: 1 = no barrier, 2 = barrier. If an object is defined as a barrier, the movement stops as soon as it impacts the object;
- Object score in the range 0-6 associated to the case or -9999 for no data.
This file requires a header line the content of which, however, does not matter. The following example defines two objects:
| ID | TYPE | SCORE |
| 2 | 2 | 3 |
| 5 | 1 | 4 |

models
Information on the models to be applied
Without the flags m and s, five parameters are required for each model:
- Model id;
- Type of model;
- Model parameters a, b and c.
Given that two models shall be used, the input could look as follows:
models=1,3,1.9,0.16,0.83,2,1,11,-9999,-9999
In this case, the model with the id of 1 would use the model of type 3, which needs all three parameters a, b and c. The model with the ID of 2 would employ the model of type 1, which only needs the parameter a. Therefore the remaining two parameters are not used. It is recommended to set unused parameters to -9999.
Five types of models are available:
| Model type | Model equation | Example reference |
| 1 | omegaT = a | Zimmermann et al. (1997) for debris flows |
| 2 | log10(tan(omegaT))=a*log_10(V)+b | Scheidegger (1973) for rock avalanches |
| 3 | Lmax=a*V^b*Z^c | Rickenmann (1999) for debris flows |
| 4 | omegaT=a*Qp^b | Huggel (2004) for lake outburst floods |
| 5 | vT≤0 | Two-parameter friction model, Perla et al. (1980) |
omegaT = angle of reach, Lmax = maximum horizontal travel distance, Z = vertical distance of motion at termination, V = volume of motion, Qp = peak discharge, vT= velocity of motion at termination.
Five additional parameters are needed with the flag s:
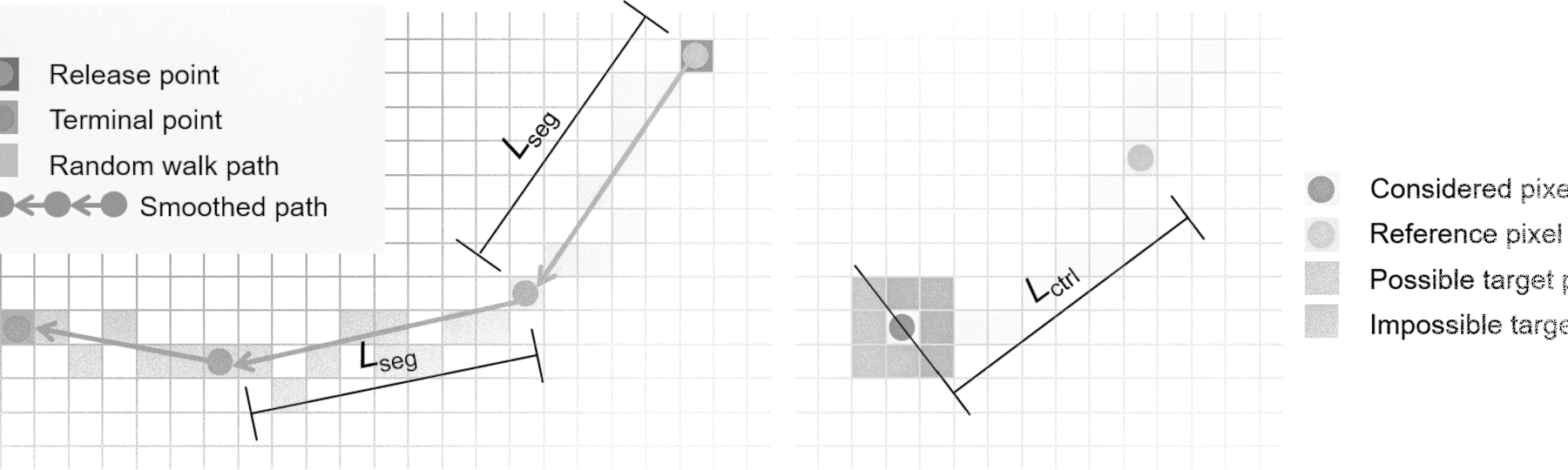
- Control length in metres Lctrl;
- Segment length in metres Lseg for the computation of the travel distance and (for model type 5) the velocity;
- Maximum runup in metres Rmax;
- Weighting factor for slope fbeta;
- Weighting factor for perpetuation of flow direction fdir.
Given that the same two models as in the above example shall be used, the input could look as follows:
models=1,3,1.9,0.16,0.83,5000,100,10,5,2,2,1,11,-9999,-9999,500,100,5,5,2
With the flag m, minima and maxima for constraining the randomization or controlled variation (option sampling) for all parameters except model id and model type have to be defined instead of single values. For controlled sampling (sampling=0), a third value denoting the number of values to be tested has to be entered for each parameter. Also for one-at-a-time sampling (sampling= lower than 0), a third value has to be entered for each parameter, denoting its initial estimate. To avoid varying a specific parameter, identical values have to be entered for the minimum and the maximum:
models=1,3,1.6,2.2,0.06,0.26,0.63,1.03,5000,5000,100,100,10,10,5,5,2,2,2,1,7,15,-9999,-9999,-9999,-9999,500,500,100,100,5,5,5,5,2,2
Zimmermann, M., Mani, P., Gamma, P.: Murganggefahr und Klimaänderung - ein GIS basierter Ansatz. NFP 31 Schlussbericht, Hochschulverlag an der ETH, Zürich, 1997.
Scheidegger, A. E. (1973): On the Prediction of the Reach and Velocity of Catastrophic Landslides. Rock Mechanics 5: 231-236.
Rickenmann, D. (1999): Empirical Relationships for Debris Flows. Natural Hazards, 19: 47-77.
Huggel, C. (2004): Assessment of Glacial Hazards based on Remote Sensing and GIS Modeling. Dissertation at the University of Zurich, Schriftenreihe Physische Geographie Glaziologie und Geomorphodynamik.
Perla, R., Cheng, T. T., McClung, D. M. (1980): A Two-Parameter Model of Snow Avalanche Motion. Journal of Glaciology 26: 197-207.

mparams
Control parameters for the random walk routing
With the flag s, only two comma-separated parameters have to be specified:
- The logarithm (base 10) of the number of random walks to be performed for each case;
- The minimum horizontal travel distance Lmin in metres. The break criteria are tested only after the motion has passed this distance. For some types of mass movements, Lmin may be set to zero, in other cases it may be useful to choose values larger than zero. This allows to start routing even if the area defined by the release coordinates would be too flat. With the flag b (back-calculation of observations), all sets of random walks with a maximum travel distance Lmax smaller than Lmin are excluded from the resulting statistics.
Without the flag s, also the five comma-separated parameters Lctrl, Lseg, Rmax, fbeta and fdir (see option models) have do be defined here, directly after log10(nwalks) and Lmin. This means that the same set of the parameters is applied to all models:
mparams=5,20,1000,100,10,5,2
With the flag m, minima and maxima for constraining the randomization or controlled variation (option sampling) have to be entered for all parameters instead of single values. For controlled sampling (sampling=0), a third value denoting the number of values to be tested has to be entered for each parameter. Also for one-at-a-time sampling (sampling= lower than 0), a third value has to be entered for each parameter, denoting its initial estimate. To avoid varying a specific parameter, identical values have to be provided for the minimum and the maximum:
mparams=5,5,20,20,200,2000,100,100,10,20,5,5,2,2

sampling
Type of parameter sampling
This parameter is mandatory along with the flags m and n. With the flag m it defines the type of sampling of the parameters given by models and mparams: integer values larger than 0 = random sampling (the default; the value given denotes the number of model runs i.e., the sample size), 0 = controlled sampling, integer values smaller than 0 = one-at-a-time sampling (the value given, if multiplied by -1, denotes the number of model runs i.e., the sample size, for each parameter). With controlled or one-at-a-time sampling, the number of model runs is applied to generate a uniform distribution between the minimum and the maximum of each parameter which is varied. This means that the actual number of model runs is the product of the sample sizes associated to all the varied parameters. With the flag n, sampling has to be a positive integer number defining the number of random subsets of the area of interest the model is applied to.

retain
Per cent of cases to be retained for evaluation
Optional floating point number defining the per cent of cases to be retained for evaluation (applicable with the flag n). The default is 20.

functype
Type of pdf to be applied
Optional integer number defining the type of probability density function to be applied along with the flags b or n. 1 = Gaussian (the default), 2 = log-normal.

backfile
Name of text file with back-calculated angles of reach and travel distances
Relative path to a text file summarizing the back-calculation of the angle of reach and the maximum travel distance of observed mass movements. This file is produced automatically with the flag b and is a mandatory input with the flag n. The file has to consist of four columns. A header line is mandatory, the columns have to be named as follows:
- ID: running id of the set of random walks;
- CASE: case id as defined by the releasefile or the releasemap;
- LMAX: maximum travel distance associated to the set of random walks;
- OMEGAT: angle of reach associated to the set of random walks.

cdffile
Name of text file with cumulative pdf
Relative path to a text file associating a cumulative probability density function to selected threshold levels of the tangent of the angle of reach. This parameter is mandatory if the flag p is active whilst the flag n is inactive. The cdffile has to consist of two columns. A header line is mandatory, the columns have to be named as follows:
- OMEGAT: the tangents of the values of the angle of reach have to be given in ascending order in the range of 0-2 at an interval of 0.001;
- CDF: cumulative density associated to each value of the angle of reach.

zonalfile
Name of text file with parameters needed to compute the zonal release probability
Relative path to a text file defining the seven parameters needed for computing the zonal release probability. zonalfile is mandatory if the option probmap is given in combination with the flag p. The file consists of a header and seven lines, each of them showing the parameter name in the first column and the parameter value in the second column.

profile
Profile along main flow path
Main flow path, given by the coordinates of an unlimited number of points along the flow path, separated by commas in the following sequence: x of first point, y of first point, x of second point and so on (all in metres). Note that the sequence of points has to strictly follow the course of the path, starting at the top and ending at the bottom, without jumping forth and back.

Results
r.randomwalk 2.1 automatically creates raster maps, graphics, and other files.
The names of all output raster maps, folders and files start with the prefix defined by the option prefix. r.randomwalk produces a set of output GRASS raster maps stored in the active mapset as well as a set of txt and tif files stored in subfolders of the folder [prefix]_results:
- Input files for model evaluation, parameter sensitivity analysis and parameter optimization with the software AIMEC are stored in the subfolder [prefix]_aimec;
- Exported ascii raster maps for use with other software packages are stored in the subfolder [prefix]_ascii;
- Text files are stored in the subfolder [prefix]_files;
- Map and diagram plots visualizing the key results are stored in the subfolder [prefix]_plots.
The subfolder [prefix]_plots, including its content, is produced only if the flag v (evaluation and visualization of model results) is active. The folder [prefix]_aimec is only produced with the flag m.

GRASS raster maps
Result raster maps stored in the active GRASS Mapset
Active GRASS Mapset
r.randomwalk produces the following output raster maps which are, however, only stored permanently with the flag k:
- [prefix]_if
Impact frequency All modes - [prefix]_iii
Impact indicator index m - [prefix]_pi
Impact probability p - [prefix]_picounter
Number of model runs affecting raster cell n+p - [prefix]_pi_stdev
Standard deviation of impact probability a+n+p - [prefix]_pic
Composite probability p+probmap - [prefix]_prz
Zonal release probability p+probmap - [prefix]_probcorr
Correction function for zonal release probability p+probmap - [prefix]_arz
Area of possible release zone p+probmap - [prefix]_prz_stdev
Standard deviation of zonal release probability p+probmap - [prefix]_iis
Impact indicator score q - [prefix]_ihis
Hazard indicator score q+scoremap - [prefix]_ris
Risk indicator score q+scoremap+impactobjects+objectscores[prefix]_velocity Flow velocity Model type 5 - [prefix]_sconn
Connectivity score c 
AIMEC input
Input files for forther processing with AIMEC
[prefix]_aimec
With the flag m, this folder contains the automatically generated input folders and files for the evaluation, parameter sensitivity analysis and optimization tool AIMEC (Fischer, 2013). In order to enable applying this tool to the results produced by r.randomwalk, the content and structure of this folder should not be manipulated manually.
Fischer, J.-T. (2013): A novel approach to evaluate and compare computational snow avalanche simulation. Natural Hazards and Earth System Sciences 13: 1655-1667.

ASCII rasters
ASCII rasters of results which can be exported for use in other software packages
[prefix]_ascii
The output raster maps are exported as ascii rasters in order to be accessible with other software packages. The naming scheme follows the one used for the GRASS raster maps.

Text files
Text files describing the simulation and its results
[prefix]_files
The following text files summarize the main parameters of the simulation:
- [prefix]_allparams.txt: This file summarizes all input and evaluation parameters (with flag m only). It is produced by merging the files [prefix]_params.txt and [prefix]_evaluation.txt (see below).
- [prefix]_averages.txt: this file is provided along with the flag p and the option probmap, or with the flag m. It summarizes the fraction of observed mass movement impact raster cells in the studied area and the average of the modelled composite probability or impact indicator index.
- [prefix]_cdf.txt: cumulative probability densities for selected threshold levels of the tangent of the angle of reach. This file is produced in the back-calculation mode (flag b) and required as input for model runs with the flag p if the flag n is not active (option cdffile).
- [prefix]_param.txt: parameter file, produced without the flag m only.
- [prefix]_params.txt: this file is produced along with the flag m only. It summarizes the input parameters for all model runs. For each columnm, the id of the model run is followed by the parameters given in the same sequence as in mparams and models.
- [prefix]_backfile.txt: result of the back-calculation of maximum travel distances and angles of reach of observed landslides (produced with flag b and needed with flag n, see option backfile).
- [prefix]_sconn.txt: connectivity score. This file is produced along with the flag c and with the use of a releasefile only. It shows for each case the total number of times random walks associated to the case hit a cell with a value larger than zero of the impactobjects raster.
- [prefix]_summary.txt: summary file of the calculation. With the flags m or n, one summary file is produced for each model run. The number of the associated model run is added at the end of the name of each file. The column headers refer to the maximum travel distance LMAX and the angle of reach OMEGAT, the suffices indicated the ids of the various models used (option models). AREA refers to the total predicted impact area in square metres.
- [prefix]_time.txt: computational time (excluding evaluation and visualization) in seconds.
- [prefix]_evaluation.txt: This file is produced along with the flag m only. It summarizes the evaluation outcomes of all model runs. Column headers: id = id of model run, nraster cells = number of affected raster cells, Lmax = maximum travel distance, omegat = angle of reach, Lim = length of modelled impact area along defined profile, Lratio = excess travel distance ratio, TP = true positive per cent, TN = true negative per cent, FP = false positive per cent, FN = false negative per cent, CSI = critical success index, HSS = Heidke skill score, AUROC = area under the ROC curve, D2PC = distance to perfect classification, FoC = factor of conservativeness, SPI = synthetic performance index. Note that some of the parameters are only given if depositmap or profile are specified. If more than one case is considered, Lmax and omegaT refer to the one with the highest case id.

Maps and graphics
Graphic representation of the main simulation results
[prefix]_plots
The set of plots produced with r.randomwalk includes:
- Colour map layouts illustrating the most relevant results. The naming scheme corresponds to the scheme introduced for the GRASS raster maps.
- If maps of the impact indicator index, the impact probability or the composite probability are included in the results, 2 ROC Plots relating each eligible result map to the observed deposition areas (option depositmap) or, for the composite probability, to the observed impact areas (option impactmap), are created: one plot includes the entire study area ([prefix]_roc.png), the second one uses a normalized TN per cent value ([prefix]_rocn.png). The area under the curve AUCROC is displayed as the key indicator for the prediction quality. With the flag n, one curve is generated for each model run. In this case, each ROC Plot includes multiple curves and the key statistics of AUCROC are shown.
- The histogram, probability density function and cumulative density function of the tangent of the angle of reach are plotted with flag b if more than one case is back-calculated. With flag n, multiple histograms, probability density and cumulative density functions are plotted (one for each model run).
- [prefix]_multval_[evaluation parameter].png: These files are produced with the flag m if the option deposit is provided, and controlled sampling with variation of two parameters is applied. A series of two-dimensional plots illustrate the sensitivity of the evaluation parameters Lratio, CSI, HSS, AUROC, D2PC, FoC, and SPI to the parameters which are varied.
Two additional text files, aucroc.txt and aucrocn.txt, are produced along with the ROC Plots. These files are stored directly in the working directory and display the prefix of the computation along with the corresponding value of AUCROC: in aucroc.txt, the value refers to the entire area of interest, while in aucrocn.txt, a normalized TN per cent value is used. These two files are not overwritten during the next execution of r.randomwalk. Instead, a new line is added each time r.randomwalk is run with the flag v. This facilitates the analysis of the AUCROC values in case of multiple executions of r.randomwalk.

Please cite this site and its content as: Mergili, M., 2014-2021. r.randomwalk - The landslide routing tool. r.randomwalk 2.1 User manual. https://www.randomwalk.org/manual.php






